重新下載正確HRV檔案,尋找HRV原始檔(5min 靜息)
- 共 246 個
- 有 5min session 的:229 個
- 沒有 5min session 的:17 個
- S11204(只有 生理曲線試作.prt)
- S112071, S112077, S112084, S112121, S112156, S112157, S112159, S112161, S112162, S112170, S112171, S112183, S112185, S112189, S112192
- S123345(只有 P5 Physiology BVP Temp…)
資料架構
- Protocol.dat 顯示 310 秒(5min)
- Session.dat 實際資料長度換算後約 294.46 秒(接近 5 分鐘)
- 這是因為 Session.dat 是 int16 且為 5 通道、取樣率 2048 Hz
- 5 通道 * 2048 Hz * 294.46s * 2 bytes ≈ 6.0MB(吻合)
- chs 檔顯示通道順序是 A=HR/BVP(血容積脈波), B=EKG(心電), C=Temp(溫度), D=Resp(呼吸), E=GSR(皮膚電導)
前處理 (對channel A & channel B)
- band-pass filter:0.5–100 Hz(去除 baseline wander 等)
- notch filter:60 Hz(去除電源干擾)
仍缺原始檔(9 人)
- S112008
- S112019
- S112070
- S112158
- S112159
- S112160
- S112171
- S112183
- S112192
Lasso Ranking
/Users/yuchi/PycharmProjects/PsyMl_ISI/.venv/bin/python /Users/yuchi/PycharmProjects/PsyMl_ISI/ML/tools/lasso_ranking.py
[模式] 允許模式
[資料] 來源=raw_data_hrv_recalc 目標=3TP 樣本=80 特徵=10
已排除群組:['ACS', 'CPT', 'EEG', 'IGT', 'ISI', 'PSQI', 'WM']
總缺值比例:45.00%
[CV結果](分數 = neg_log_loss,越大越好 → log_loss 越小)
lambda_min:C = 0.145202 | lambda = 6.88696
lambda_2SE:C = 0.0316228 | lambda = 31.6228
[選入變項(2SE)] 以 |係數| 排序(前 30)
(無變項被選入;可放寬正則或檢查特徵)
[對照] lambda_min 非零變項數:0,lambda_2SE 非零變項數:0
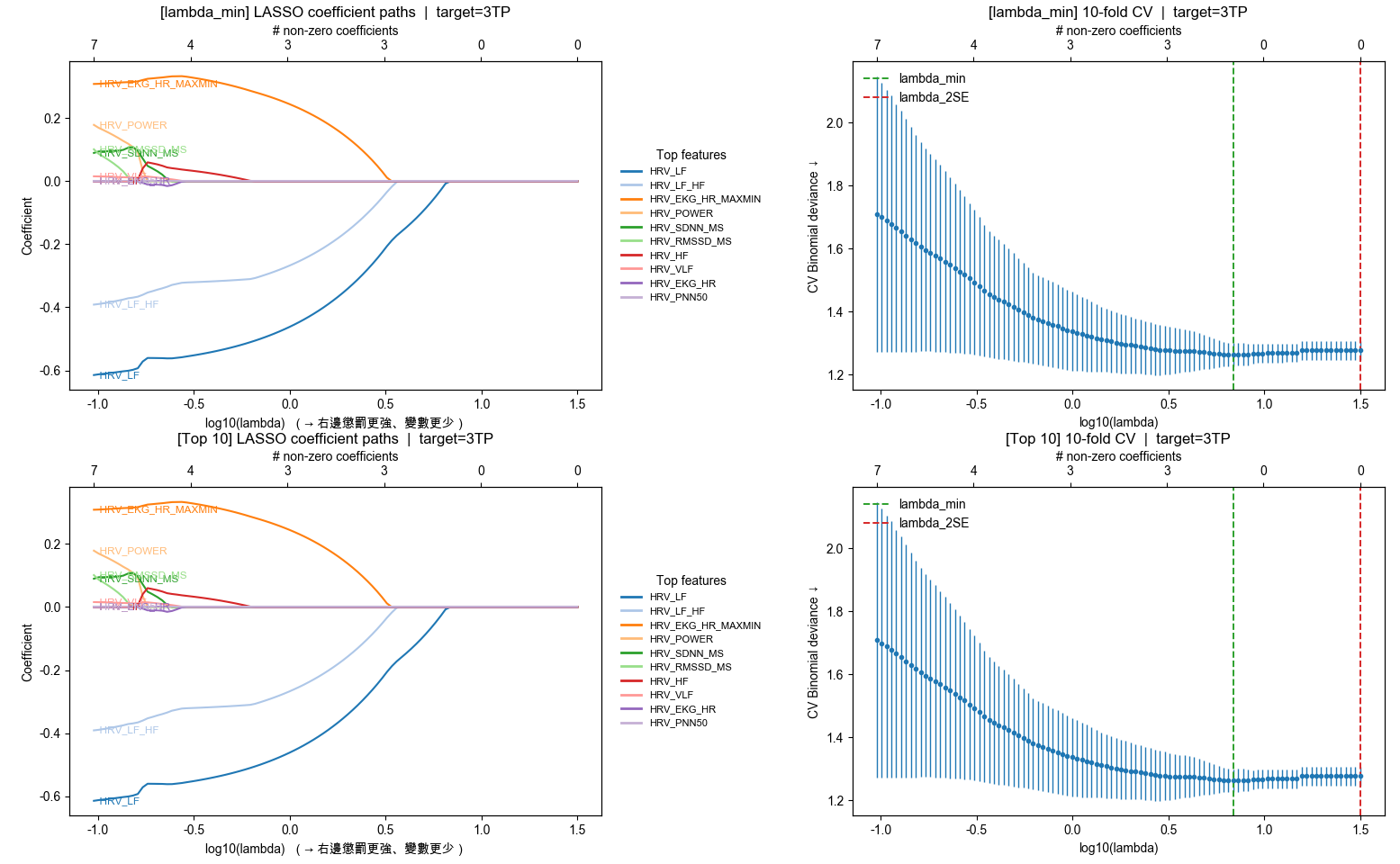
[Top 10(路徑峰值)] 不綁定單一 C
HRV_LF
HRV_LF_HF
HRV_EKG_HR_MAXMIN
HRV_POWER
HRV_SDNN_MS
HRV_RMSSD_MS
HRV_HF
HRV_VLF
HRV_EKG_HR
HRV_PNN50
ML
單次數據
BAI_T1,BDI_T1
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=2
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
LogisticRegression 0.955 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
NaiveBayes 0.955 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
MLP 0.938 0.788 0.684 0.929 0.873 0.960 0.800 0.685 0.841 3.800 5.000
SVM 0.931 0.733 0.688 0.786 0.862 0.893 0.833 0.599 0.818 3.200 5.600
KNN 0.929 0.750 0.900 0.643 0.906 0.853 0.967 0.677 0.864 2.000 6.800
RandomForest 0.926 0.741 0.769 0.714 0.885 0.871 0.900 0.627 0.841 2.600 6.200
XGBoost 0.926 0.714 0.714 0.714 0.867 0.867 0.867 0.581 0.818 2.800 6.000
DecisionTree 0.755 0.667 0.692 0.643 0.852 0.839 0.867 0.520 0.795 2.600 6.200
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum
LogisticRegression 28 2 3 11
NaiveBayes 28 2 3 11
MLP 24 6 1 13
SVM 25 5 3 11
KNN 29 1 5 9
RandomForest 27 3 4 10
XGBoost 26 4 4 10
DecisionTree 26 4 5 9
=== LOSO (Leave-One-Subject-Out) — WEIGHTED metrics ===
BAI_T1,BDI_T1,HRV_LF,HRV_LF_HF
/Users/yuchi/PycharmProjects/PsyMl_ISI/.venv/bin/python /Users/yuchi/PycharmProjects/PsyMl_ISI/ML/ml_benchmark_modular.py
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=4
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1', 'HRV_LF', 'HRV_LF_HF']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
LogisticRegression 0.964 0.769 0.833 0.714 0.903 0.875 0.933 0.677 0.864 2.400 6.400
NaiveBayes 0.957 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
MLP 0.945 0.609 0.778 0.500 0.862 0.800 0.933 0.500 0.795 1.800 7.000
SVM 0.943 0.720 0.818 0.643 0.889 0.848 0.933 0.620 0.841 2.200 6.600
RandomForest 0.936 0.714 0.714 0.714 0.867 0.867 0.867 0.581 0.818 2.800 6.000
KNN 0.918 0.750 0.900 0.643 0.906 0.853 0.967 0.677 0.864 2.000 6.800
XGBoost 0.905 0.759 0.733 0.786 0.881 0.897 0.867 0.641 0.841 3.000 5.800
DecisionTree 0.824 0.769 0.833 0.714 0.903 0.875 0.933 0.677 0.864 2.400 6.400
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum
LogisticRegression 28 2 4 10
NaiveBayes 28 2 3 11
MLP 28 2 7 7
SVM 28 2 5 9
RandomForest 26 4 4 10
KNN 29 1 5 9
XGBoost 26 4 3 11
DecisionTree 28 2 4 10
多輪數據重複運算(確認泛化能力)
BAI_T1,BDI_T1
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=2
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1']
[CV] Repeated Stratified 5-fold x 100, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Repeated Stratified 5-fold x 100 CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
NaiveBayes NaN 0.825 0.857 0.794 0.922 0.907 0.938 0.748 0.892 2.594 6.206
LogisticRegression NaN 0.808 0.832 0.786 0.914 0.903 0.926 0.723 0.881 2.648 6.152
MLP NaN 0.785 0.669 0.949 0.866 0.971 0.781 0.684 0.835 3.972 4.828
SVM NaN 0.759 0.729 0.791 0.880 0.899 0.863 0.641 0.840 3.038 5.762
KNN NaN 0.753 0.856 0.671 0.902 0.861 0.947 0.666 0.860 2.196 6.604
XGBoost NaN 0.727 0.712 0.743 0.869 0.878 0.860 0.596 0.823 2.920 5.880
RandomForest NaN 0.714 0.709 0.719 0.865 0.868 0.862 0.579 0.817 2.838 5.962
DecisionTree NaN 0.630 0.661 0.602 0.839 0.822 0.856 0.470 0.775 2.550 6.250
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum
NaiveBayes 2815 185 288 1112
LogisticRegression 2777 223 299 1101
MLP 2343 657 71 1329
SVM 2589 411 292 1108
KNN 2842 158 460 940
XGBoost 2580 420 360 1040
RandomForest 2587 413 394 1006
DecisionTree 2568 432 557 843
BAI_T1,BDI_T1,HRV_LF,HRV_LF_HF
/Users/yuchi/PycharmProjects/PsyMl_ISI/.venv/bin/python /Users/yuchi/PycharmProjects/PsyMl_ISI/ML/ml_benchmark_modular.py
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=4
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1', 'HRV_LF', 'HRV_LF_HF']
[CV] Repeated Stratified 5-fold x 100, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Repeated Stratified 5-fold x 100 CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
NaiveBayes NaN 0.801 0.843 0.764 0.914 0.894 0.934 0.717 0.880 2.536 6.264
LogisticRegression NaN 0.787 0.843 0.738 0.909 0.884 0.936 0.700 0.873 2.452 6.348
KNN NaN 0.743 0.897 0.634 0.904 0.850 0.966 0.669 0.860 1.978 6.822
RandomForest NaN 0.718 0.766 0.675 0.880 0.856 0.904 0.600 0.831 2.466 6.334
SVM NaN 0.715 0.801 0.646 0.885 0.849 0.925 0.609 0.836 2.260 6.540
XGBoost NaN 0.714 0.734 0.696 0.872 0.861 0.882 0.587 0.823 2.654 6.146
DecisionTree NaN 0.643 0.660 0.628 0.839 0.830 0.849 0.483 0.779 2.664 6.136
MLP NaN 0.633 0.603 0.667 0.815 0.836 0.795 0.450 0.754 3.100 5.700
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum
NaiveBayes 2801 199 331 1069
LogisticRegression 2807 193 367 1033
KNN 2898 102 513 887
RandomForest 2712 288 455 945
SVM 2775 225 495 905
XGBoost 2647 353 426 974
DecisionTree 2547 453 521 879
MLP 2384 616 466 934
偷看答案方式:
BAI_T1, BDI_T1
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=2
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
LogisticRegression 0.955 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
NaiveBayes 0.955 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
MLP 0.938 0.788 0.684 0.929 0.873 0.960 0.800 0.685 0.841 3.800 5.000
SVM 0.931 0.733 0.688 0.786 0.862 0.893 0.833 0.599 0.818 3.200 5.600
KNN 0.929 0.750 0.900 0.643 0.906 0.853 0.967 0.677 0.864 2.000 6.800
RandomForest 0.926 0.741 0.769 0.714 0.885 0.871 0.900 0.627 0.841 2.600 6.200
XGBoost 0.926 0.714 0.714 0.714 0.867 0.867 0.867 0.581 0.818 2.800 6.000
DecisionTree 0.755 0.667 0.692 0.643 0.852 0.839 0.867 0.520 0.795 2.600 6.200
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum FP_FN_IDS
LogisticRegression 28 2 3 11 [S112119, S112194 | S112036, S112002, S112169]
NaiveBayes 28 2 3 11 [S112119, S112194 | S112036, S112002, S112169]
MLP 24 6 1 13 [S112030, S112104, S112119, S112194, S112012, S112047 | S112115]
SVM 25 5 3 11 [S112030, S112104, S112119, S112194, S112047 | S112036, S112002, S112169]
KNN 29 1 5 9 [S112194 | S112036, S112039, S112002, S112169, S112029]
RandomForest 27 3 4 10 [S112194, S112012, S112047 | S112036, S112039, S112002, S112169]
XGBoost 26 4 4 10 [S112119, S112194, S112012, S112047 | S112036, S112039, S112002, S112169]
DecisionTree 26 4 5 9 [S112119, S112194, S112012, S112047 | S112023, S112036, S112039, S112002, S112169]
HRV_LF, HRV_LF_HF
/Users/yuchi/PycharmProjects/PsyMl_ISI/.venv/bin/python /Users/yuchi/PycharmProjects/PsyMl_ISI/ML/ml_benchmark_modular.py
[資料] 來源=raw_data_hrv_recalc 目標=3TP 列數=44 特徵數=4
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1', 'HRV_LF', 'HRV_LF_HF']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 30 68.2
1 14 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
LogisticRegression 0.962 0.769 0.833 0.714 0.903 0.875 0.933 0.677 0.864 2.400 6.400
NaiveBayes 0.960 0.815 0.846 0.786 0.918 0.903 0.933 0.734 0.886 2.600 6.200
RandomForest 0.936 0.714 0.714 0.714 0.867 0.867 0.867 0.581 0.818 2.800 6.000
SVM 0.926 0.769 0.833 0.714 0.903 0.875 0.933 0.677 0.864 2.400 6.400
KNN 0.911 0.696 0.889 0.571 0.892 0.829 0.967 0.621 0.841 1.800 7.000
XGBoost 0.905 0.759 0.733 0.786 0.881 0.897 0.867 0.641 0.841 3.000 5.800
DecisionTree 0.824 0.769 0.833 0.714 0.903 0.875 0.933 0.677 0.864 2.400 6.400
MLP 0.733 0.485 0.421 0.571 0.691 0.760 0.633 0.193 0.614 3.800 5.000
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum FP_FN_IDS
LogisticRegression 28 2 4 10 [S112119, S112194 | S112036, S112002, S112169, S112029]
NaiveBayes 28 2 3 11 [S112119, S112194 | S112036, S112002, S112169]
RandomForest 26 4 4 10 [S112119, S112194, S112012, S112047 | S112036, S112039, S112002, S112169]
SVM 28 2 4 10 [S112119, S112194 | S112036, S112002, S112169, S112029]
KNN 29 1 6 8 [S112194 | S112023, S112036, S112039, S112002, S112169, S112029]
XGBoost 26 4 3 11 [S112119, S112194, S112012, S112047 | S112036, S112039, S112002]
DecisionTree 28 2 4 10 [S112119, S112194 | S112036, S112039, S112002, S112029]
MLP 19 11 6 8 [S112044, S112176, S112045, S112077, S112173, S112119, S112194, S112026, S112043, S112047, S112177 | S112055, S112036, S112115, S112039, S112002, S112169]
- 從偷看答案的方式可以看出,模型用
BAI_T1,BDI_T1原本就分得不錯,只有單獨幾個一直分不出來(S112119, S112194 | S112036, S112002, S112169) 👉 分不出來的可能是異常值 - 加入
HRV_LF,HRV_LF_HF之後,原本分不出來的還是分不出來,而且反而產生噪音讓更多樣本分不出(S112029)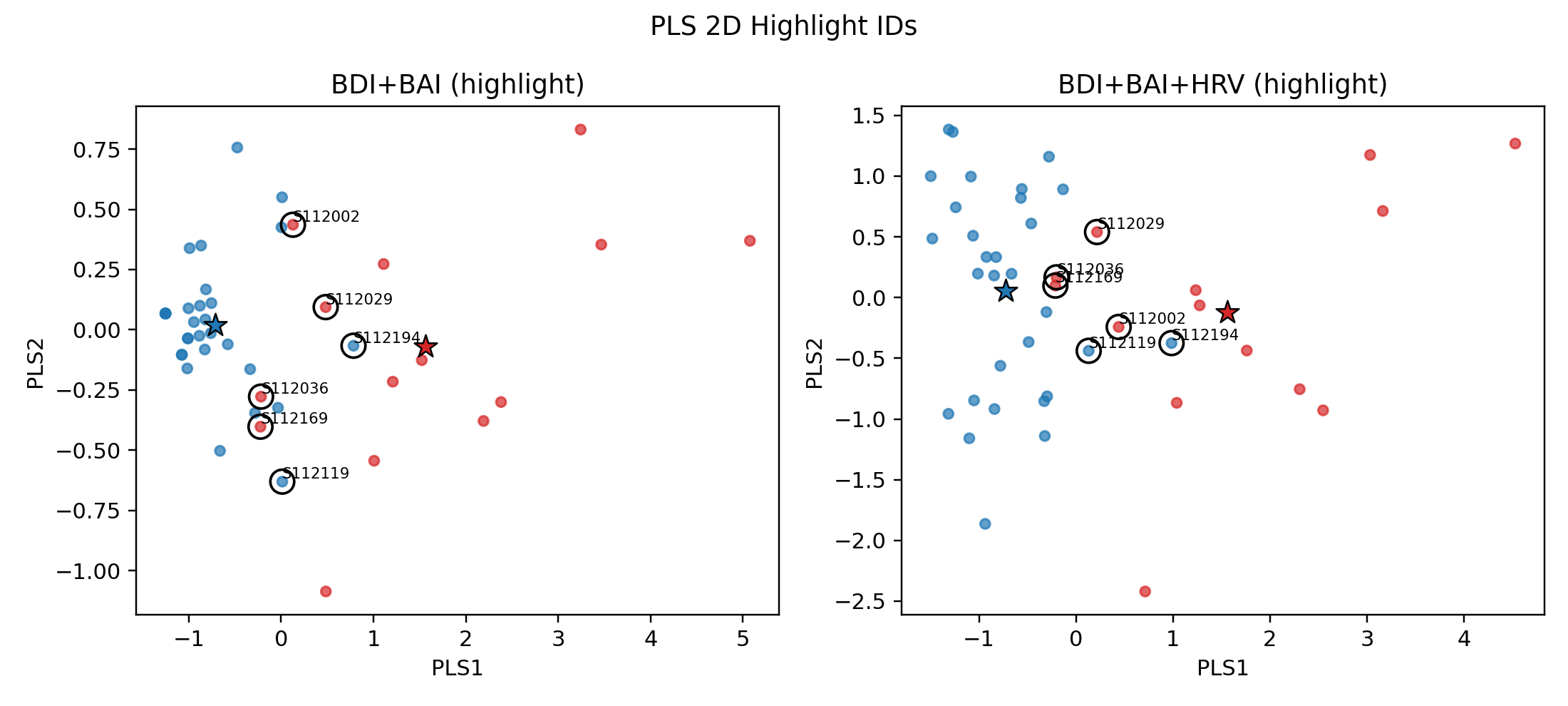
- 註:將算錯的樣本用PLS_2D呈現(黑色圈圈處),發現加入HRV後發而讓紅色(失眠群)往藍色星星(非失眠群)靠近 👉 會讓模型更難分辨
PLS監督分析 (PLS_DA回歸)
-
為什麼明明
HRV應該可以幫助預測失眠,但事實上卻讓數據變差:- 目前樣本數下(少樣本),HRV 屬於低 SNR 的增量特徵,導致模型方差上升與決策邊界不穩定 👉 HRV有提供分類幫助,但自身噪音or異常值的影響大於幫助影響
- 因此 AUC 可能略升,但閾值下的 F1/混淆矩陣反而變差
-
PLS觀察方法:PLS可以用2D/3D呈現分類的樣本中心點(紅色/藍色星星),樣本離該類別星星越近,代表ML模型可分性越高(越好分類)
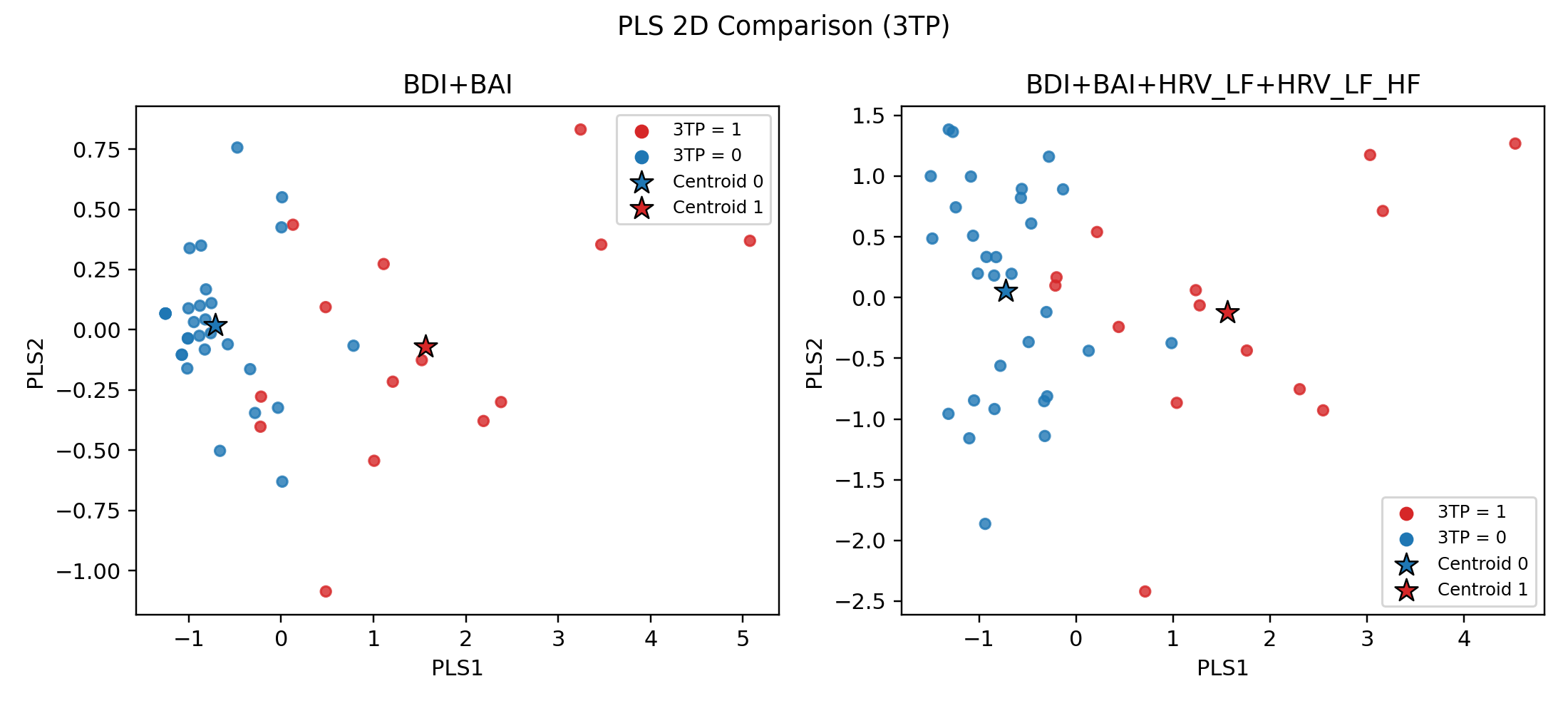
- 藍色樣本(健康樣本)原本很好預測,加入HRV之後反而把樣本間距拉開
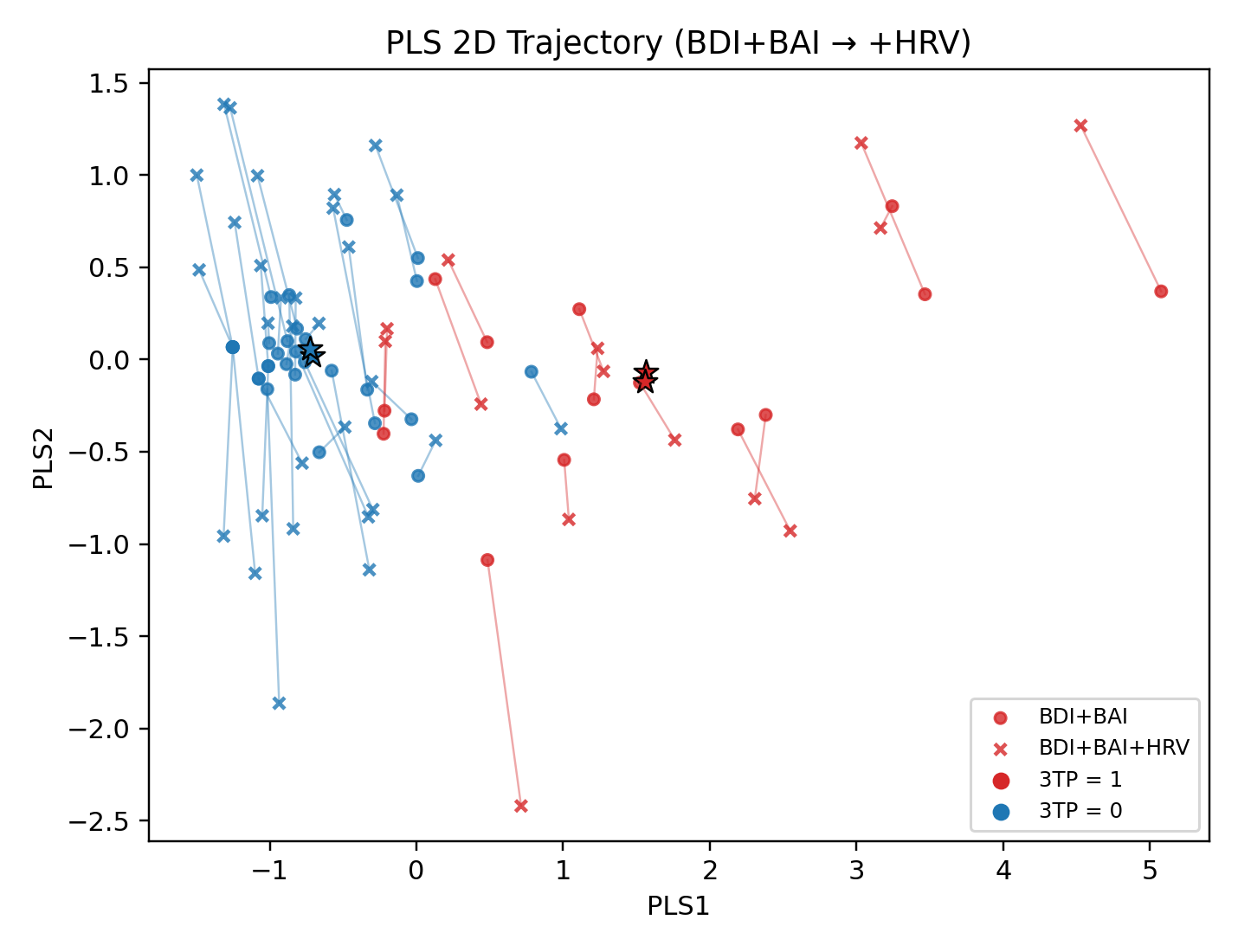
- 樣本變化軌跡,X表示加入HRV之後樣本的位置(實心為原始位置)
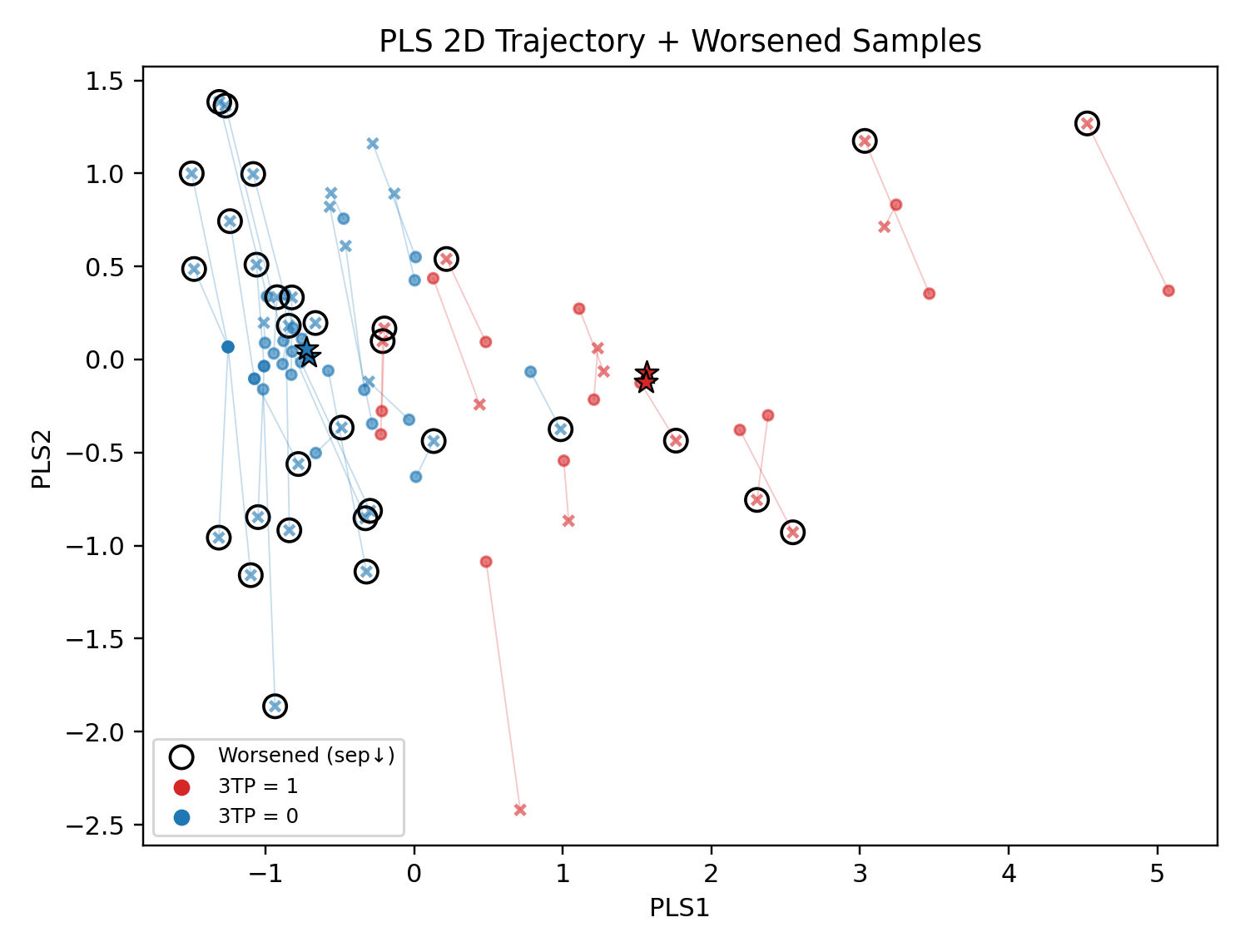
- 加入HRV之後導致更難預測的樣本,用黑色框出(sep下降)