📊 資料對稱性統計
總體樣本概況
| 指標 | 樣本數 | 說明 |
|---|---|---|
| 總樣本數 | 331 | 資料庫中所有受試者 |
| T1 完整 | 326 | ISI_T1 + BDI_T1 + BAI_T1 非空 |
| T1 + T2 完整 | 259 | ISI_T1, ISI_T2, BDI_T1, BDI_T2, BAI_T1, BAI_T2 非空 |
| T1 + T2 + T3 完整 | 156 | 所有測時點和基線特徵都非空 |
目標變數與可用樣本
| 目標變數 | 樣本數 | 比例 | 說明 |
|---|---|---|---|
| 3TP (目標) | 97 | 29.3% | 用於 3 測時點預測 |
| 2TP (目標) | 191 | 57.7% | 用於 2 測時點預測 |
生物標記可用性
| 生物標記類型 | 樣本數 | 完整性 | 說明 |
|---|---|---|---|
| EEG (FAA) | 267 | 80.7% | EEG_FAA_REL_ALPHA_F4F3 非空 |
| HRV (RMSSD) | 267 | 80.7% | HRV_RMSSD_MS 非空 |
| PAC (MVL) | 273 | 82.5% | isi_raw_data_transformer_pac_mvl 配對 |
| PAC (MI) | 273 | 82.5% | isi_raw_data_transformer_pac_mi 配對 |
基線特徵與附加特徵組合
| 組合類型 | 樣本數 | 完整度 | 說明 |
|---|---|---|---|
| BDI + BAI (基線) | 326 | 98.5% | 基線特徵最高完整度 |
| BDI + BAI + EEG_FAA | 263 | 79.5% | 加入 EEG 特徵 |
| BDI + BAI + EEG_FAA + 3TP | 66 | 19.9% | 用於本次分析的完整樣本 |
| BDI + BAI + EEG_FAA + 2TP | 139 | 42.0% | 2TP 目標的完整樣本 |
🔑 關鍵發現
- 對稱最好的測時點: T1 有 326 個完整樣本 (98.5%)
- 主要瓶頸: 3TP 目標變數稀缺,只有 97 個樣本 → 與完整特徵組合交集後只剩 66 個
- 生物標記覆蓋率: PAC > EEG ≈ HRV (都在 80-82%)
- 建議: 可考慮用 2TP (191 個) 或擴大 T1+T2 樣本 (259 個) 進行分析
[資料] 來源=isi_raw_data_transformer 目標=3TP 列數=66 特徵數=2
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 45 68.2
1 21 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC AUC_overall F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
NaiveBayes 0.869 0.869 0.684 0.765 0.619 0.872 0.837 0.911 0.565 0.818 3.400 9.800
LogisticRegression 0.868 0.868 0.651 0.636 0.667 0.831 0.841 0.822 0.483 0.773 4.400 8.800
SVM 0.807 0.807 0.638 0.577 0.714 0.800 0.850 0.756 0.448 0.742 5.200 8.000
KNN 0.789 0.789 0.526 0.588 0.476 0.809 0.776 0.844 0.342 0.727 3.400 9.800
RandomForest 0.784 0.784 0.649 0.750 0.571 0.863 0.820 0.911 0.524 0.803 3.200 10.000
MLP 0.746 0.746 0.579 0.647 0.524 0.830 0.796 0.867 0.416 0.758 3.400 9.800
DecisionTree 0.695 0.695 0.558 0.545 0.571 0.787 0.795 0.778 0.345 0.712 4.400 8.800
XGBoost 0.690 0.690 0.600 0.632 0.571 0.826 0.809 0.844 0.428 0.758 3.800 9.400
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum FP_FN_IDS
NaiveBayes 41 4 8 13 [S112019, S112240, S112271, S112194 | S112008, S112036, S112257, S112203, S112029, S112169, S112183, S112268]
LogisticRegression 37 8 7 14 [S112019, S112047, S112240, S112271, S112194, S112030, S112119, S112159 | S112008, S112036, S112257, S112203, S112222, S112169, S112183]
SVM 34 11 6 15 [S112184, S112201, S112019, S112047, S112240, S112271, S112194, S112030, S112119, S112159, S112168 | S112008, S112257, S112203, S112222, S112169, S112183]
KNN 38 7 11 10 [S112019, S112047, S112240, S112271, S112194, S112119, S112159 | S112008, S112036, S112214, S112039, S112257, S112203, S112222, S112029, S112169, S112183, S112268]
RandomForest 41 4 9 12 [S112019, S112047, S112240, S112271 | S112008, S112086, S112039, S112203, S112222, S112029, S112169, S112183, S112268]
MLP 39 6 10 11 [S112184, S112019, S112042, S112047, S112240, S112271 | S112008, S112086, S112214, S112039, S112203, S112222, S112029, S112169, S112268, S112023]
DecisionTree 35 10 9 12 [S112019, S112026, S112186, S112207, S112215, S112247, S112047, S112240, S112271, S112030 | S112008, S112086, S112214, S112039, S112203, S112222, S112029, S112183, S112268]
XGBoost 38 7 9 12 [S112019, S112215, S112047, S112240, S112271, S112159, S112168 | S112008, S112036, S112086, S112039, S112203, S112222, S112029, S112183, S112268]
[資料] 來源=isi_raw_data_transformer 目標=3TP 列數=66 特徵數=3
[特徵] 使用欄位(前 15):['BDI_T1', 'BAI_T1', 'EEG_FAA_REL_ALPHA_F4F3']
[CV] Stratified 5-fold, seed=42 | class_weight=balanced
[聚合] K-fold = weighted | LOSO = weighted
[Leakage check] Class balance
count percent%
3TP
0 45 68.2
1 21 31.8
[Leakage check] 未發現與目標 |r| ≥ 0.95 的欄位。
=== Basic ML Benchmark (Stratified 5-fold CV) ===
model AUC AUC_overall F1_pos(=1) Prec_pos Rec_pos F1_neg(=0) Prec_neg Rec_neg MCC Accuracy Pred1_mean Pred0_mean
NaiveBayes 0.901 0.901 0.703 0.812 0.619 0.884 0.840 0.933 0.600 0.833 3.200 10.000
LogisticRegression 0.863 0.863 0.667 0.667 0.667 0.844 0.844 0.844 0.511 0.788 4.200 9.000
MLP 0.848 0.848 0.732 0.750 0.714 0.879 0.870 0.889 0.611 0.833 4.000 9.200
KNN 0.842 0.842 0.600 0.632 0.571 0.826 0.809 0.844 0.428 0.758 3.800 9.400
RandomForest 0.838 0.838 0.579 0.647 0.524 0.830 0.796 0.867 0.416 0.758 3.400 9.800
SVM 0.833 0.833 0.711 0.667 0.762 0.851 0.881 0.822 0.566 0.803 4.800 8.400
XGBoost 0.782 0.782 0.564 0.611 0.524 0.817 0.792 0.844 0.385 0.742 3.600 9.600
DecisionTree 0.695 0.695 0.579 0.647 0.524 0.830 0.796 0.867 0.416 0.758 3.400 9.800
--- Aggregated Confusion Matrix Sums (across all folds' test parts) ---
model TN_sum FP_sum FN_sum TP_sum FP_FN_IDS
NaiveBayes 42 3 8 13 [S112019, S112240, S112271 | S112008, S112036, S112257, S112203, S112029, S112169, S112183, S112268]
LogisticRegression 38 7 7 14 [S112019, S112026, S112047, S112240, S112271, S112194, S112159 | S112008, S112036, S112257, S112203, S112222, S112169, S112183]
MLP 40 5 6 15 [S112019, S112186, S112047, S112240, S112271 | S112008, S112039, S112257, S112203, S112029, S112023]
KNN 38 7 9 12 [S112201, S112019, S112047, S112240, S112271, S112030, S112159 | S112008, S112214, S112039, S112257, S112203, S112029, S112169, S112183, S112268]
RandomForest 39 6 10 11 [S112019, S112247, S112047, S112240, S112271, S112159 | S112008, S112039, S112257, S112075, S112203, S112222, S112029, S112169, S112183, S112268]
SVM 37 8 5 16 [S112201, S112019, S112215, S112047, S112240, S112271, S112030, S112159 | S112008, S112203, S112222, S112169, S112183]
XGBoost 38 7 10 11 [S112201, S112019, S112247, S112047, S112240, S112271, S112159 | S112008, S112036, S112086, S112039, S112257, S112075, S112203, S112222, S112029, S112268]
DecisionTree 39 6 10 11 [S112105, S112215, S112047, S112135, S112240, S112271 | S112008, S112086, S112039, S112075, S112203, S112222, S112029, S112183, S112268, S112023]
Lasso Ranking
[警告] isi_raw_data_transformer_pac_mvl 裡 NEW_NUMBER 有重複(保留第一筆):['S112116', 'S112158']
[PAC JOIN] isi_raw_data_transformer_pac_mvl: 67/97 個樣本匹配
[警告] isi_raw_data_transformer_pac_mi 裡 NEW_NUMBER 有重複(保留第一筆):['S112116', 'S112158']
[PAC JOIN] isi_raw_data_transformer_pac_mi: 67/97 個樣本匹配
[樣本模式] 排除模式 - 使用所有受試者(後續可能依缺值規則排除)
[模式] 排除模式
[篩選] 必須完整欄位數=2973 | 受試者 97 → 66
[資料] 來源=isi_raw_data_transformer 目標=3TP 樣本=66 特徵=3007
已排除群組:['ACS', 'CPT', 'IGT', 'ISI', 'PSQI', 'WM']
總缺值比例:0.06%
使用模型:LASSO
補值策略:自動補中位數
受試者篩選:啟用(要求特定欄位組完整)
[CV結果](分數 = neg_log_loss,越大越好 → log_loss 越小)
lambda_min:C = 0.137733 | lambda = 7.26043
lambda_1SE:C = 0.0764579 | lambda = 13.0791
[選入變項(1SE)] 以 |係數| 排序(前 30)
coef abs_coef data_ratio
BDI_T1 0.40035 0.40035 1.0
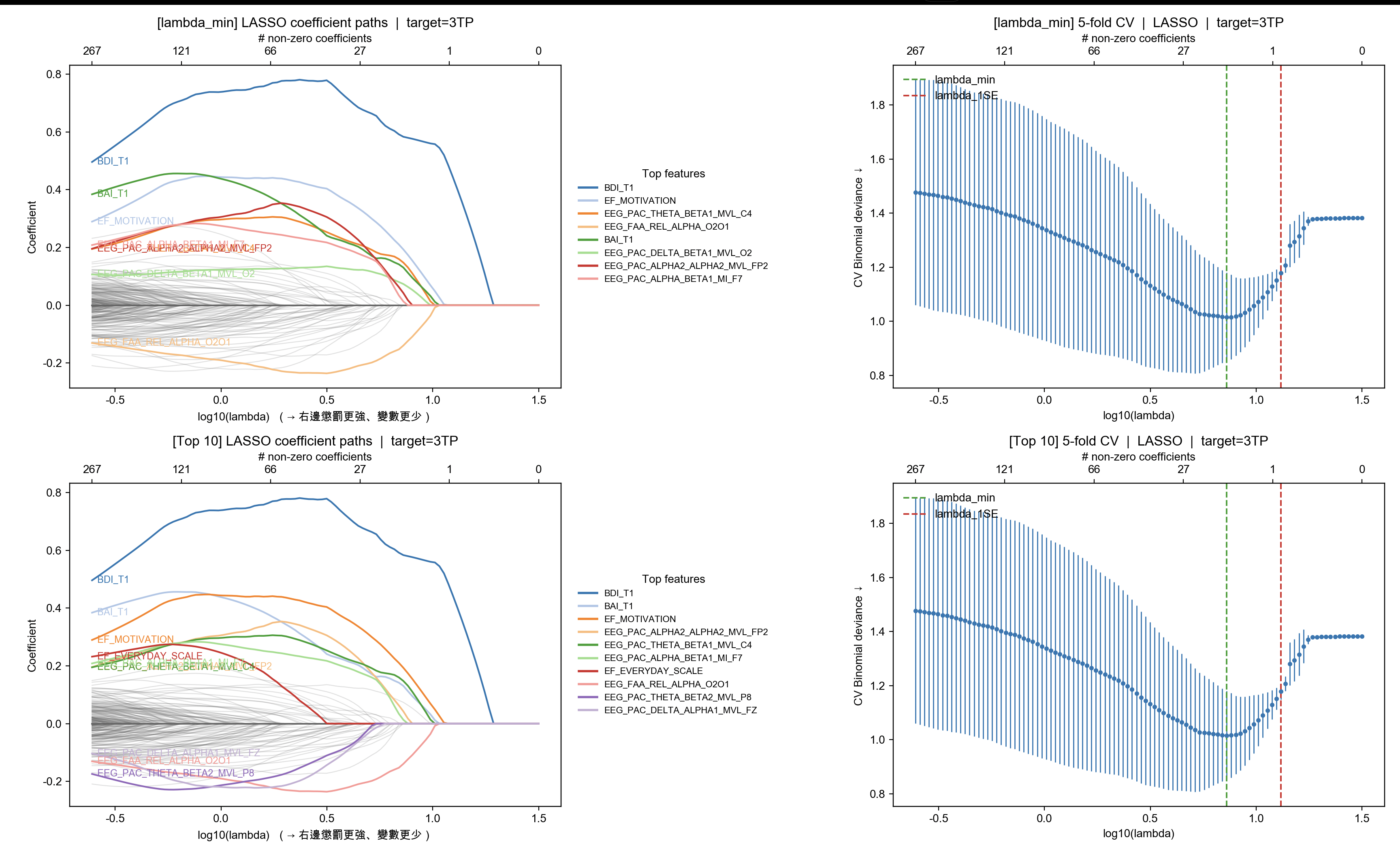
[對照] lambda_min 非零變項數:8,lambda_1SE 非零變項數:1
[選入變項(lambda_min)] 以 |係數| 排序(前 30)
coef abs_coef data_ratio
BDI_T1 0.583288 0.583288 1.000000
EF_MOTIVATION 0.193174 0.193174 0.954545
EEG_PAC_THETA_BETA1_MVL_C4 0.147609 0.147609 1.000000
EEG_FAA_REL_ALPHA_O2O1 -0.139352 0.139352 1.000000
BAI_T1 0.130506 0.130506 1.000000
EEG_PAC_DELTA_BETA1_MVL_O2 0.073537 0.073537 1.000000
EEG_PAC_ALPHA2_ALPHA2_MVL_FP2 0.042265 0.042265 1.000000
EEG_PAC_ALPHA_BETA1_MI_F7 0.013178 0.013178 1.000000
────────────────────────────────────────────────────────────────────────────────
Baseline 各模型 AUC
────────────────────────────────────────────────────────────────────────────────
DecisionTree F1=0.4751 AUC=0.6535
KNN F1=0.5087 AUC=0.7949
LogisticRegression F1=0.6042 AUC=0.8589
MLP F1=0.4907 AUC=0.8177
NaiveBayes F1=0.6129 AUC=0.8875
RandomForest F1=0.4828 AUC=0.8142
SVM F1=0.6264 AUC=0.8226
XGBoost F1=0.3848 AUC=0.7808
────────────────────────────────────────────────────────────────────────────────
Top 30 附加特徵 (mean_delta_AUC 排序)
────────────────────────────────────────────────────────────────────────────────
feature_combination mean_delta positive_ratio delta_DecisionTree delta_KNN delta_LogisticRegression delta_MLP delta_NaiveBayes delta_RandomForest delta_SVM delta_XGBoost
rank
1 +EEG_PAC_THETA_BETA2_MVL_P4 0.0599 1.00 0.0387 0.1515 0.0391 0.0532 0.0474 0.0462 0.0960 0.0074
2 +EEG_PAC_THETA_BETA1_MVL_F8 0.0599 0.88 0.0475 0.1297 -0.0238 0.1212 0.0234 0.0544 0.0819 0.0450
3 +EEG_PWR_ABS_GAMMA2_CZ 0.0540 0.88 0.0364 0.1505 0.0206 0.0808 -0.0292 0.0610 0.0514 0.0603
4 +EEG_PWR_ABS_GAMMA_CZ 0.0495 0.75 -0.0230 0.1506 0.0268 0.0653 -0.0006 0.0548 0.0514 0.0706
5 +EEG_PAC_ALPHA1_BETA_MVL_CZ 0.0495 0.88 0.0618 0.0985 -0.0079 0.0929 0.0175 0.0206 0.0347 0.0777
6 +EEG_PAC_ALPHA1_BETA_MVL_F4 0.0491 0.88 0.1518 0.0796 -0.0171 0.0016 0.0143 0.0365 0.0294 0.0967
7 +EEG_PWR_ABS_GAMMA1_CZ 0.0473 0.88 -0.0253 0.1292 0.0204 0.0593 0.0121 0.0659 0.0514 0.0654
8 +EEG_PWR_ABS_GAMMA2_C3 0.0473 1.00 0.0588 0.0567 0.0173 0.0802 0.0016 0.0327 0.0419 0.0891
9 +EEG_PAC_DELTA_ALPHA_MI_C3 0.0468 1.00 0.1049 0.1027 0.0169 0.0385 0.0079 0.0445 0.0222 0.0369
10 +EEG_PWR_ABS_HGAMMA_CZ 0.0467 0.88 0.0722 0.1430 0.0238 0.0077 -0.0861 0.0718 0.0514 0.0901
11 +EEG_PAC_ALPHA_ALPHA1_MI_FP2 0.0460 0.75 0.0357 0.0672 -0.0101 0.0539 -0.0157 0.0707 0.0663 0.0997
12 +EEG_PAC_ALPHA1_ALPHA_MI_C3 0.0444 0.75 0.1955 0.0413 0.0327 -0.0097 -0.0437 0.0354 0.0163 0.0870
13 +EEG_PAC_DELTA_BETA1_MVL_O2 0.0430 1.00 0.0341 0.0870 0.0391 0.0212 0.0204 0.0265 0.0575 0.0577
14 +EEG_PAC_THETA_ALPHA_MVL_C4 0.0425 0.88 0.0373 0.0839 0.0548 0.0750 0.0179 0.0365 0.0458 -0.0115
15 +EEG_PAC_THETA_ALPHA2_MVL_O1 0.0402 0.75 0.0089 0.1169 0.0173 0.0885 -0.0048 0.0452 0.0696 -0.0202
16 +EEG_FAA_REL_ALPHA_F4F3 0.0402 1.00 0.0454 0.0900 0.0111 0.0554 0.0298 0.0573 0.0321 0.0008
17 +EEG_PAC_ALPHA1_BETA_MI_CZ 0.0393 0.88 0.0491 0.0569 -0.0109 0.0976 0.0111 0.0438 0.0254 0.0413
18 +EEG_PAC_ALPHA_ALPHA_MI_C3 0.0391 0.75 -0.0112 0.1102 -0.0077 0.0222 0.0550 0.0237 0.0550 0.0656
19 +EEG_PAC_ALPHA_BETA_MVL_F4 0.0386 0.88 0.0677 0.0982 -0.0107 0.0502 0.0365 0.0373 0.0236 0.0057
20 +EEG_FAA_REL_GAMMA_C4C3 0.0386 1.00 0.0595 0.0750 0.0113 0.0702 0.0175 0.0283 0.0202 0.0266
21 +EEG_PAC_ALPHA2_GAMMA2_MI_O2 0.0385 0.75 -0.0001 0.0991 0.0615 0.0720 0.0327 0.0141 -0.0087 0.0370
22 +EEG_PAC_DELTA_GAMMA_MVL_T7 0.0377 0.88 0.0722 0.0956 0.0264 0.0256 -0.0079 0.0339 0.0470 0.0087
23 +EEG_PWR_ABS_GAMMA_C3 0.0376 1.00 0.0246 0.0849 0.0236 0.0579 0.0048 0.0209 0.0389 0.0452
24 +EEG_PAC_ALPHA1_BETA_MI_F4 0.0375 0.88 0.1019 0.0591 -0.0105 0.0063 0.0149 0.0390 0.0238 0.0657
25 +EEG_PAC_ALPHA2_ALPHA1_MVL_T7 0.0375 0.75 -0.0133 0.0766 -0.0012 0.0440 0.0355 0.0400 0.0446 0.0736
26 +EEG_PAC_DELTA_ALPHA2_MVL_P3 0.0369 0.88 0.0262 0.0786 0.0242 0.0575 -0.0050 0.0352 0.0260 0.0526
27 +EEG_PAC_ALPHA_GAMMA1_MI_CZ 0.0360 1.00 0.0863 0.0352 0.0143 0.0274 0.0085 0.0298 0.0226 0.0639
28 +EEG_FAA_REL_GAMMA2_C4C3 0.0355 0.88 0.0944 0.0751 0.0113 0.0546 0.0143 0.0317 -0.0050 0.0076
29 +EEG_PAC_ALPHA2_GAMMA2_MI_P7 0.0351 0.75 0.1301 0.0674 -0.0204 0.0500 -0.0238 0.0236 0.0115 0.0422
30 +EEG_FAA_REL_GAMMA2_FP2FP1 0.0351 0.88 0.0142 0.0750 0.0081 0.0284 -0.0111 0.0385 0.0482 0.0792
────────────────────────────────────────────────────────────────────────────────
Top 10 by positive_ratio(對最多模型有幫助)
────────────────────────────────────────────────────────────────────────────────
feature_combination mean_delta positive_ratio delta_DecisionTree delta_KNN delta_LogisticRegression delta_MLP delta_NaiveBayes delta_RandomForest delta_SVM delta_XGBoost
rank
1 +EEG_PAC_THETA_BETA2_MVL_P4 0.0599 1.0 0.0387 0.1515 0.0391 0.0532 0.0474 0.0462 0.0960 0.0074
8 +EEG_PWR_ABS_GAMMA2_C3 0.0473 1.0 0.0588 0.0567 0.0173 0.0802 0.0016 0.0327 0.0419 0.0891
9 +EEG_PAC_DELTA_ALPHA_MI_C3 0.0468 1.0 0.1049 0.1027 0.0169 0.0385 0.0079 0.0445 0.0222 0.0369
13 +EEG_PAC_DELTA_BETA1_MVL_O2 0.0430 1.0 0.0341 0.0870 0.0391 0.0212 0.0204 0.0265 0.0575 0.0577
16 +EEG_FAA_REL_ALPHA_F4F3 0.0402 1.0 0.0454 0.0900 0.0111 0.0554 0.0298 0.0573 0.0321 0.0008
20 +EEG_FAA_REL_GAMMA_C4C3 0.0386 1.0 0.0595 0.0750 0.0113 0.0702 0.0175 0.0283 0.0202 0.0266
23 +EEG_PWR_ABS_GAMMA_C3 0.0376 1.0 0.0246 0.0849 0.0236 0.0579 0.0048 0.0209 0.0389 0.0452
27 +EEG_PAC_ALPHA_GAMMA1_MI_CZ 0.0360 1.0 0.0863 0.0352 0.0143 0.0274 0.0085 0.0298 0.0226 0.0639
33 +EEG_PAC_ALPHA2_BETA3_MVL_T7 0.0342 1.0 0.0253 0.0484 0.0327 0.0280 0.0044 0.0785 0.0210 0.0351
37 +EEG_PAC_THETA_GAMMA2_MI_T7 0.0330 1.0 0.0350 0.0920 0.0363 0.0201 0.0329 0.0232 0.0179 0.0065
================================================================================
完成。總耗時:768.4s (12.8 min)
================================================================================
暴力拆解解釋:
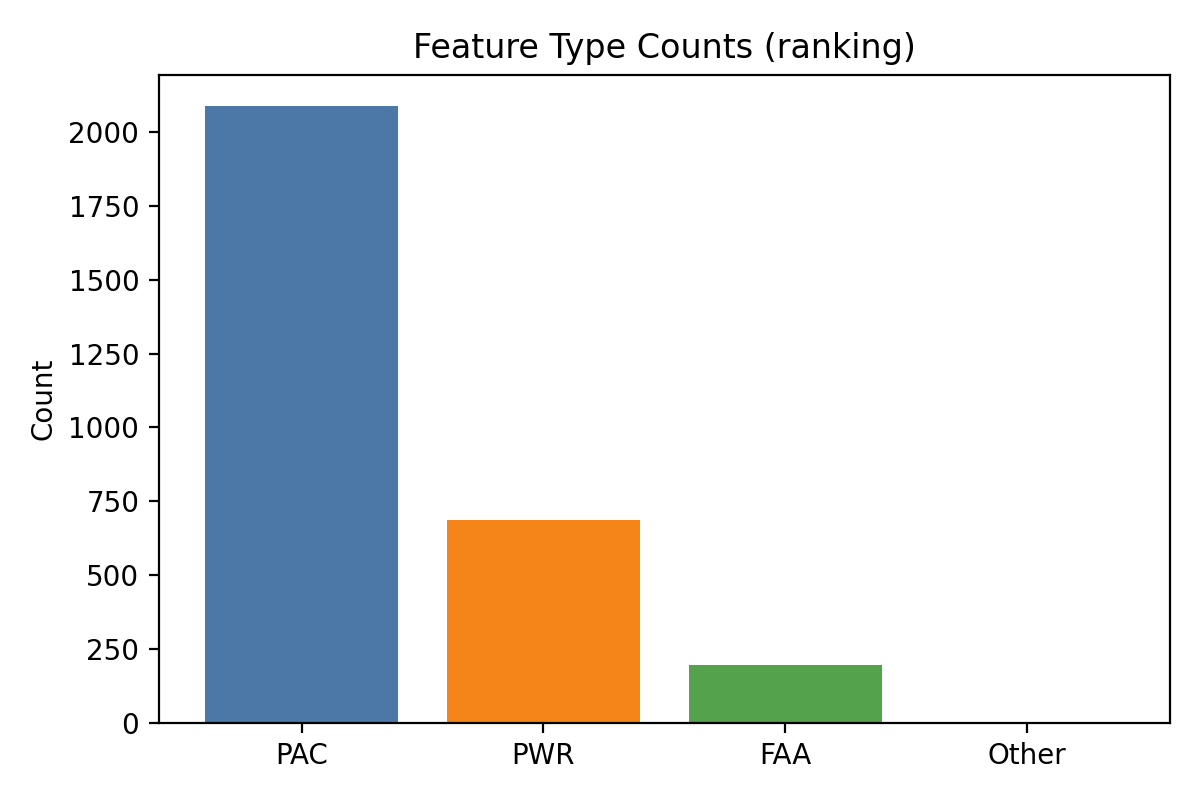
- 比例
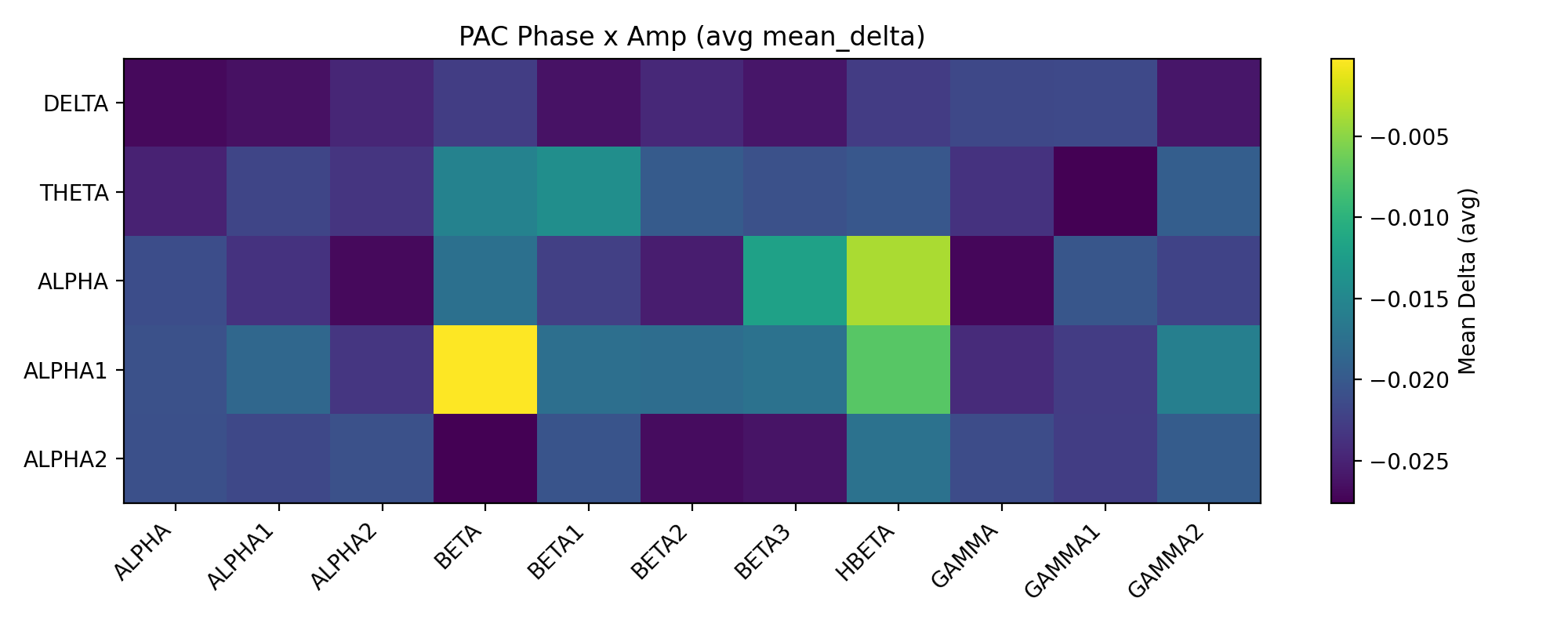
- 耦合熱圖
- 考慮到所有通道
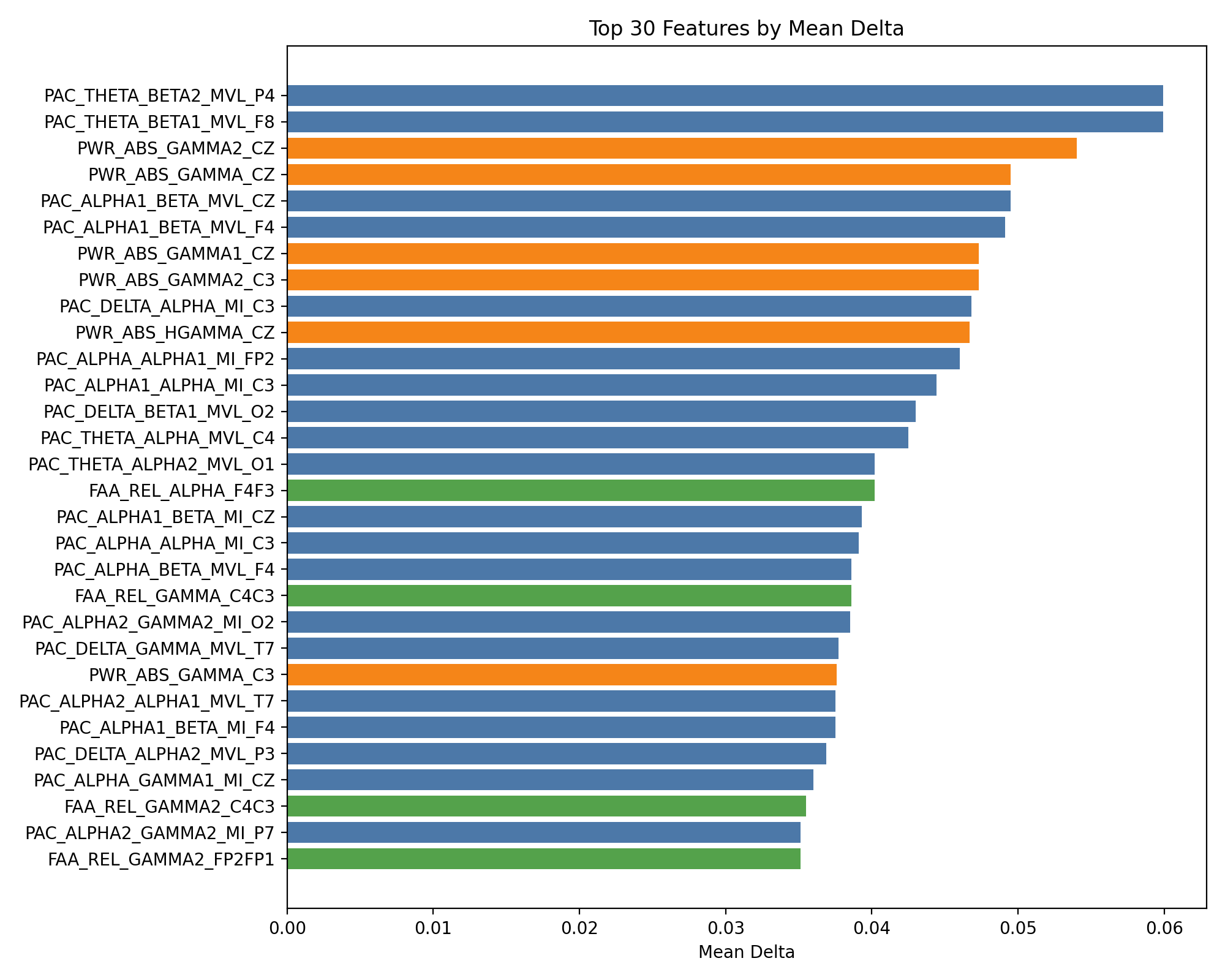
增加更多 PWR 相加
📍 EEG_PWR 點位相加統計
資料庫中 EEG_PWR 欄位採用兩層結構:
- 單點位 (Single Electrodes): C3, C4, F3, F4, F7, F8, FP1, FP2, O1, O2, P3, P4, CZ, FZ, PZ, T3, T4, T5, T6
- 相加組合 (Averaged Pairs): 標記為
_AVG結尾,例如C3C4_AVG,F3F4_AVG
🔄 相加組合統計總表
| 頻段 | 頻率範圍 | 相加組合數 | PWR 類型 | 典型相加位置 |
|---|---|---|---|---|
| Delta | 0.5-3 Hz | 19×2 = 38 | ABS, REL | BRAIN_AVG, C3C4_AVG, F3F4_AVG, F7F8_AVG, FP_AVG, FZCZPZ_AVG, O1O2_AVG, P3P4_AVG, T3T4_AVG, T5T6_AVG |
| Theta | 4-7 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Alpha1 | 8-10 Hz | 18×2 = 36 | ABS, REL | 同上 (少 F3F4_AVG) |
| Alpha2 | 11-13 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Alpha | 8-13 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Beta1 | 13-20 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Beta2 | 20-30 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Beta3 | 30-40 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Beta | 13-40 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Gamma1 | 30-50 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Gamma2 | 40-80 Hz | 19×2 = 38 | ABS, REL | 同上 |
| Gamma | 30-80 Hz | 19×2 = 38 | ABS, REL | 同上 |
| High Beta | 20-40 Hz | 19×2 = 38 | ABS, REL | 同上 |
| High Gamma | 60-100 Hz | 19×2 = 38 | ABS, REL | 同上 |
總計: 14 個頻段 × (平均 19-18 個相加組合) × 2 (ABS/REL) ≈ 500+ 個 PWR 相加欄位
📊 相加位置解析
| 位置代碼 | 說明 | 包含的電極 | 功能區域 |
|---|---|---|---|
| BRAIN_AVG | 全腦平均 | 所有電極 | 全局腦活動 |
| C3C4_AVG | 中央相加 | C3 + C4 | 中央皮質 (Motor area) |
| F3F4_AVG | 前額相加 | F3 + F4 | 背外側前額皮質 (DLPFC) |
| F7F8_AVG | 顳額相加 | F7 + F8 | 側顳皮質 |
| FP_AVG | 前極相加 | FP1 + FP2 | 前額極 (PFC) |
| FZCZPZ_AVG | 中線相加 | FZ + CZ + PZ | 中線皮質 |
| O1O2_AVG | 枕葉相加 | O1 + O2 | 視覺皮質 |
| P3P4_AVG | 頂葉相加 | P3 + P4 | 頂葉皮質 |
| T3T4_AVG | 顳葉相加 | T3 + T4 | 側顳皮質 (lower) |
| T5T6_AVG | 後顳葉相加 | T5 + T6 | 後側顳皮質 |
💡 設計理念
- 左右對稱相加: 大多數相加組合為左右半球對稱的電極對,便於檢測半球間的功能整合
- 雙重量化: 同時提供 ABS (絕對功率) 和 REL (相對功率),便於歸一化比較
- 跨頻段覆蓋: 從低頻 (Delta) 到高頻 (High Gamma),涵蓋認知和神經生理的全頻譜
🎯 常用組合組合建議
推薦用於睡眠/放鬆分析:
- Delta + Theta 的
BRAIN_AVG(低頻全腦) - Alpha 的
FZCZPZ_AVG(中線放鬆)
推薦用於認知/執行功能:
- Beta1 + Beta2 的
F3F4_AVG(前額執行控制) - Gamma 的
C3C4_AVG(中央認知處理)
推薦用於情緒/社交:
- Alpha1/2 的
F7F8_AVG(顳葉社交處理) - Theta 的
O1O2_AVG(後顳葉整合)